Summary
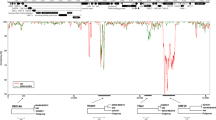
The genome of adenovirus (AV) types 1, 2, and 5 is known to be very variable as evidenced by the great number of genome types described (36, 61, and 35 for AV1, 2, and 5 respectively). Physical maps were constructed for nearly all of the restriction or R-variants by biochemical methods and by adapting restriction fragments of defined molecular weight. The alterations found with seven DNA restriction endonucleases were mapped on the genome. Altered restriction sites, found for the genome types of each of the three serotypes, appeared to be randomly distributed over the genome, although mutations seemed to occur preferentially on distinct sites of the genome. Many strains, especially those of AV5, were genomically much more related with one another than with the prototype. This finding is compatible with the interpretation that the related strains were derived from an unknown common ancestor.
Similar content being viewed by others
References
Adrian T, Bastian B, Wagner V (1989) Restriction site mapping of adenovirus types 9 and 15 and genome types of intermediate 15/H9. Intervirology 30: 169–176
Adrian T, Sassinek J, Wigand R (1990) Genome type analysis of 480 isolates of adenovirus types 1, 2, and 5. Arch Virol 112: 235–248
Adrian T, Wadell G, Hierholzer JC, Wigand R (1986) DNA restriction analysis of adenovirus prototypes 1 to 41. Arch Virol 91: 277–290
Adrian T, Wolf U (1989) Adenovirus 6 genome types: mapping of restriction site alterations on the genome. Res Virol 140: 545–550
Aird F, King JJ, Younghusband HB (1983) Identification of a new strain of adenovirus type 2 by restriction endonuclease analysis. Gene 22: 133–134
Boursnell MEG, Mautner V (1981) Recombinations in adenoviruses: crossover sites in intertypic recombinants are located in regions of homology. Virology 112: 198–209
Broker TR (1980) Nucleotide sequence and analysis of group-C adenoviruses. In: Tooze J (ed) DNA tumor viruses. Molecular biology of tumor viruses, 2nd ed, part 2. Cold Spring Harbour Laboratory Press, New York, pp 923–935
Chroboczek J, Bieber F, Jacrot B (1992) The sequence of the genome of adenovirus type 5 and its comparison with the genome of adenovirus type 2. Virology 186: 280–285
Fife KH, Ashley R, Corey L (1985) Isolation and characterisation of six new genome types of human adenovirus types 1 and 2. J Clin Microbiol 21: 20–23
Fox JP, Hall CE, Cooney MK (1977) The Seattle virus watch. VII. Observations of adenovirus infections. Am J Epidemiol 105: 362–386
Green M, Mackey JK, Wold WSM, Rigden P (1979) Thirty-one human adenovirus serotypes (Ad1–31) from five groups (A–E) based upon DNA genome homologies. Virology 93: 481–492
Horwitz MS (1990) Adenoviruses. In: Fields BN, Knipe DM (eds) Virology, 2nd ed. Raven Press, New York, pp 1723–1740
Kaier B, Wigand R (1986) Antigenic homogeneity of adenovirus types 1, 2, 5, and 6. J Med Virol 18: 283–287
Kinloch R, Mackay N, Mautner V (1984) Adenovirus hexon. Sequence comparison of subgroup C serotypes 2 and 5. J Biol Chem 259: 6431–6436
Legerski RJ, Hodnett JL, Gray HB (1978). Extracellular nucleases of pseudomonas Bal31 III. Use of the double-strand deoxyribonuclease activity as the basis for the mapping of a convenient method for the mapping of fragments of DNA produced by cleavage with restriction enzymes. Nucleic Acids Res 5: 1445–1464
Niel C, D'Halluin JC (1984) Restriction maps of human adenovirus types 2, 3, and 5 for BstEII, MluI, NruI, and SfiI endonucleases. Gene 31: 305–308
Rigby PWJ, Dieckman M, Rhodes C, Berg P (1977) Labelling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol 113: 237–251
Roberts RJ, Akusjärvi G, Aleström P, Gelinas RE, Gingeras TR, Sciaky D, Pettersson U (1986) A consensus sequence for the adenovirus-2 genome. In: Doerfler W (ed) Adenovirus DNA. The viral genome and its expression. Martinus Nijhoff, Boston Dordrecht Lancaster, pp 1–51
Schmitz H, Wigand R, Heinrich W (1983) Worldwide epidemiology of human adenovirus infections. Am J Epidemiol 117: 455–466
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98: 503–517
Strohl WA, Schlesinger RW (1965) Quantitative studies of natural and experimental adenovirus infections of human cells. I. Characterisation of viral multiplication in fibroblasts derived by long-term culture from tonsils. Virology 6: 99–207
Webb DH, Shields AF, Fife KH (1987) Genomic variation of adenovirus type 5 isolates recovered from bone marrow transplant recipients. J Clin Microbiol 25: 305–308
Wigand R, Adrian T (1986) Classification and epidemiology of adenoviruses. In: Doerfler W (ed) Adenovirus DNA. The viral genome and its expression. Martinus Nijhoff, Boston Dordrecht Lancaster, pp 409–441
Wigand R, Bartha A, Dreizin RS, Esche H, Ginsberg HS, Green M, Hierholzer JC, Kalter SS, McFerran JB, Petterson U, Russell WC, Wadell G (1982) Adenoviridae: second report. Intervirology 18: 169–176
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Adrian, T. Genome polymorphism of human adenoviruses of subgenus C. Archives of Virology 141, 1021–1031 (1996). https://doi.org/10.1007/BF01718606
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01718606