Abstract
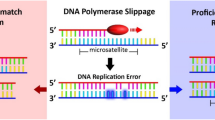
Microsatellite instability (MSI) originates from the systematic accumulation of uncorrected deletion/insertion in repetitive DNA tracts in cancer cells with a deficient mismatch repair system. Among colorectal cancers, the MSI signature identifies hereditary cases arising in patients with germline mutations in hMLH1, hMSH2, PMS2 and a fraction of those with hMSH6 mutations, as well as sporadic cancers with epigenetic hMLH1 promoter hypermethylation. Considering the specific pathogenesis, pathological features, natural history and response to 5-fluoro-uracil-based chemotherapy of the MSI cancers, confusion about the genetic markers for MSI recognition seems surprising. In this clinically relevant field, an agreement has not been reached concerning the use of di- or mononucleotide markers for MSI assessment. The Revised Bethesda Guidelines still recommend a panel of markers consisting of mono- and dinucleotides, despite being questioned whether it is congruous to continue to use dinucleotide markers for MSI identification. In any event, no single marker is accurate enough for MSI testing, and an awareness of their pros and cons is required for proper interpretation of results. In recent years, several papers have reported different prevalence of MSI in unrelated series, largely depending on the detection and classification method, suggesting that MSI test interpretation also requires the understanding of the phenomenon rather than simply the crude satisfaction of panel recommendations. Inaccuracies can otherwise lead to under- or overdiagnosis and inaccurate disease classification, which always have a negative impact on the clinical practice of medicine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Aaltonen L, Peltomaki P, Leach F, Sistonen P, Pylkkanen L, Mecklin J et al. (1993). Clues to the pathogenesis of familial colorectal cancer. Science 260: 812–816.
Aaltonen LA, Salovaara R, Kristo P, Canzian F, Hemminki A, Peltomaki P et al. (1998). Incidence of hereditary nonpolyposis colorectal cancer and the feasibility of molecular screening for the disease. N Engl J Med 338: 1481–1487.
Ahuja N, Mohan AL, Li Q, Stolker JM, Herman JG, Hamilton SR et al. (1997). Association between CpG island methylation and microsatellite instability in colorectal cancer. Cancer Res 57: 3370–3374.
Bocker T, Diermann J, Friedl W, Gebert J, Holinski-Feder E, Karner-Hanusch J et al. (1997). Microsatellite instability analysis: a multicenter study for reliability and quality control. Cancer Res 57: 4739–4743.
Boland C, Goel A . (2005). Somatic evolution of cancer cells. Semin Cancer Biol 15: 436–450.
Boland CR . (2007). Clinical uses of microsatellite instability testing in colorectal cancer: an ongoing challenge. J Clin Oncol 25: 754–756.
Boland CR, Thibodeau SN, Hamilton SR, Sidransky D, Eshleman JR, Burt RW et al. (1998). A national cancer institute workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 58: 5248–5257.
Carethers JM, Smith EJ, Behling CA, Nguyen L, Tajima A, Doctolero RT et al. (2004). Use of 5-fluorouracil and survival in patients with microsatellite-unstable colorectal cancer. Gastroenterology 126: 394–401.
Cawkwell L, Li D, Lewis F, Martin I, Dixon M, Quirke P . (1995). Microsatellite instability in colorectal cancer: improved assessment using fluorescent polymerase chain reaction. Gastroenterology 109: 465–471.
de la Chapelle A . (2003). Microsatellite instability. N Engl J Med 349: 209–210.
Dietmaier W, Wallinger S, Bocker T, Kullmann F, Fishel R, Ruschoff J . (1997). Diagnostic microsatellite instability: definition and correlation with mismatch repair protein expression. Cancer Res 57: 4749–4756.
Fishel R, Lescoe M, Rao M, Copeland N, Jenkins N, Garber J et al. (1993). The human mutator gene homolog MSH2 and its association with hereditary nonpolyposis colon cancer. Cell 75: 1027–1038.
Giuffre G, Muller A, Brodegger T, Bocker-Edmonston T, Gebert J, Kloor M et al. (2005). Microsatellite analysis of hereditary nonpolyposis colorectal cancer-associated colorectal adenomas by laser-assisted microdissection: correlation with mismatch repair protein expression provides new insights in early steps of tumorigenesis. J Mol Diagn 7: 160–170.
Goel A, Nagasaka T, Arnold C, Inoue T, Hamilton C, Niedzwiecki D et al. (2007). The CpG island methylator phenotype and chromosomal instability are inversely correlated in sporadic colorectal cancer. Gastroenterology 132: 127–138.
Gonzalez-Garcia I, Moreno V, Navarro M, Marti-Rague J, Marcuello E, Benasco C et al. (2000). Standardized approach for microsatellite instability detection in colorectal carcinomas. J Natl Cancer Inst 92: 544–549.
Grady W . (2004). Genomic instability and colon cancer. Cancer Metastasis Rev 23: 11–27.
Gruber S . (2006). New developments in Lynch syndrome (hereditary nonpolyposis colorectal cancer) and mismatch repair gene testing. Gastroenterology 130: 577–587.
Gryfe R, Kim H, Hsieh ETK, Aronson MD, Holowaty EJ, Bull SB et al. (2000). Tumor microsatellite instability and clinical outcome in young patients with colorectal cancer. N Engl J Med 342: 69–77.
Hamilton SR, Liu B, Parsons RE, Papadopoulos N, Jen J, Powell SM et al. (1995). The molecular basis of Turcot's syndrome. N Engl J Med 332: 839–847.
Hampel H, Frankel WL, Martin E, Arnold M, Khanduja K, Kuebler P et al. (2005). Screening for the Lynch syndrome (hereditary nonpolyposis colorectal cancer). N Engl J Med 352: 1851–1860.
Hemminki A, Peltomaki P, Mecklin J, Jarvinen H, Salovaara R, Nystrom-Lahti M et al. (1994). Loss of the wild type MLH1 gene is a feature of hereditary nonpolyposis colorectal cancer. Nat Genet 8: 405–410.
Herman JG, Umar A, Polyak K, Graff JR, Ahuja N, Issa J-PJ et al. (1998). Incidence and functional consequences of hMLH1 promoter hypermethylation in colorectal carcinoma. Proc Natl Acad Sci USA 95: 6870–6875.
Hoang J-M, Cottu PH, Thuille B, Salmon RJ, Thomas G, Hamelin R . (1997). BAT-26, an indicator of the replication error phenotype in colorectal cancers and cell lines. Cancer Res 57: 300–303.
Iino H, Jass J, Simms L, Young J, Leggett B, Ajioka Y et al. (1999). DNA microsatellite instability in hyperplastic polyps, serrated adenomas, and mixed polyps: a mild mutator pathway for colorectal cancer? J Clin Pathol 52: 5–9.
Ionov Y, Peinado M, Malkhosyan S, Shibata D, Perucho M . (1993). Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature 363: 558–561.
Jenkins MA, Hayashi S, O'Shea AM, Burgart LJ, Smyrk TC, Shimizu D et al. (2007). Pathology features in Bethesda guidelines predict colorectal cancer microsatellite instability: a population-based study. Gastroenterology 133: 48–56.
Jiricny J . (2006). The multifaceted mismatch-repair system. Nat Rev Mol Cell Biol 7: 335–346.
Kane MF, Loda M, Gaida GM, Lipman J, Mishra R, Goldman H et al. (1997). Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch repair-defective human tumor cell lines. Cancer Res 57: 808–811.
Kim GP, Colangelo LH, Wieand HS, Paik S, Kirsch IR, Wolmark N et al. (2007). Prognostic and predictive roles of high-degree microsatellite instability in colon cancer: a National Cancer Institute-National Surgical Adjuvant Breast and Bowel Project Collaborative Study. J Clin Oncol 25: 767–772.
Kinzler K, Vogelstein B . (1996). Lessons from hereditary colorectal cancer. Cell 87: 159–170.
Koi M, Umar A, Chauhan DP, Cherian SP, Carethers JM, Kunkei TA et al. (1994). Human chromosome 3 corrects mismatch repair deficiency and microsatellite instability and reduces N-methyl-N′-nitro-N-nitrosoguanidine tolerance in colon tumor cells with homozygous hMLH1 mutation. Cancer Res 54: 4308–4312.
Laghi L, Bianchi P, Roncalli M, Malesci A . (2004). Re: revised Bethesda guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst 96: 1402a–1403a.
Laiho P, Launonen V, Lahermo P, Esteller M, Guo M, Herman JG et al. (2002). Low-level microsatellite instability in most colorectal carcinomas. Cancer Res 62: 1166–1170.
Liu B, Parsons R, Papadopoulos N, Nicolaides N, Lynch H, Watson P et al. (1996). Analysis of mismatch repair genes in hereditary non-polyposis colorectal cancer patients. Nat Med 2: 169–174.
Lothe RA, Peltomaki P, Meling GI, Aaltonen LA, Nystrom-Lahti M, Pylkkanen L et al. (1993). Genomic instability in colorectal cancer: relationship to clinicopathological variables and family history. Cancer Res 53: 5849–5852.
Loukola A, Eklin K, Laiho P, Salovaara R, Kristo P, Jarvinen H et al. (2001). Microsatellite marker analysis in screening for hereditary nonpolyposis colorectal cancer (HNPCC). Cancer Res 61: 4545–4549.
Maddox J . (1993). Competition and the death of science. Nature 363: 667.
Malesci A, Laghi L, Bianchi P, Delconte G, Randolph A, Torri V et al. (2007). Reduced likelihood of metastases in patients with microsatellite-unstable colorectal cancer. Clin Cancer Res 13: 3831–3839.
Malkhosyan S, Rampino N, Yamamoto H, Perucho M . (1996). Frameshift mutator mutations. Nature 382: 499–500.
Markowitz S, Wang J, Myeroff L, Parsons R, Sun L, Lutterbaugh J et al. (1995). Inactivation of the type II TGF-β receptor in colon cancer cells with microsatellite instability. Science 268: 1336–1338.
Miyaki M, Konishi M, Tanaka K, Kikuchi-Yanoshita R, Muraoka M, Yasuno M et al. (1997). Germline mutation of MSH6 as the cause of hereditary nonpolyposis colorectal cancer. Nat Genet 17: 271–272.
Moslein G, Tester D, Lindor N, Honchel R, Cunningham J, French A et al. (1996). Microsatellite instability and mutation analysis of hMSH2 and hMLH1 in patients with sporadic, familial and hereditary colorectal cancer. Hum Mol Genet 5: 1245–1252.
Muller A, Beckmann C, Westphal G, Bocker Edmonston T, Friedrichs N, Dietmaier W et al. (2006). Prevalence of the mismatch-repair-deficient phenotype in colonic adenomas arising in HNPCC patients: results of a 5-year follow-up study. Int J Colorectal Dis 21: 632–641.
NCBI. (2007). OnlineMendelian Inheritance in Man (OMIM: http://www.ncbi.nlm.nih.gov/sites/entrez?db=OMIM). Accession # hMSH2, 690309; hMLH1, *1204536; PMS2, +600259; hMSH6, +600678.
Nicolaides N, Papadopoulos N, Liu B, Wei Y, Carter K, Ruben S et al. (1994). Mutations of two PMS homologues in hereditary nonpolyposis colon cancer. Nature 371: 75–80.
Ohmiya N, Matsumoto S, Yamamoto H, Baranovskaya S, Malkhosyan S, Perucho M . (2001). Germline and somatic mutations in hMSH6 and hMSH3 in gastrointestinal cancers of the microsatellite mutator phenotype. Gene 272: 301–313.
Papadopoulos N, Nicolaides N, Liu B, Parsons R, Lengauer C, Palombo F et al. (1995). Mutations of GTBP in genetically unstable cells. Science 268: 1915–1917.
Papadopoulos N, Nicolaides N, Wei Y, Ruben S, Carter K, Rosen C et al. (1994). Mutation of a mutL homolog in hereditary colon cancer. Science 263: 1625–1629.
Parsons R, Li G, Longley M, Fang W, Papadopoulos N, Jen J et al. (1993). Hypermutability and mismatch repair deficiency in RER+ tumor cells. Cell 75: 1227–1236.
Pastrello C, Baglioni S, Tibiletti M, Papi L, Fornasarig M, Morabito A et al. (2006). Stability of BAT26 in tumours of hereditary nonpolyposis colorectal cancer patients with MSH2 intragenic deletion. Eur J Hum Genet 14: 63–68.
Peltomaki P, Aaltonen L, Sistonen P, Pylkkanen L, Mecklin J, Jarvinen H et al. (1993). Genetic mapping of a locus predisposing to human colorectal cancer. Science 260: 810–812.
Perucho M, Boland CR, Thibodeau SN, Hamilton SR, Sidransky D, Eshleman JR et al. (1999). Correspondence re: C. R. Boland et al. A National Cancer Institute workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res., 58: 5248–5257, 1998. Cancer Res 59: 249–256.
Pyatt R, Chadwick RB, Johnson CK, Adebamowo C, de la Chapelle A, Prior TW . (1999). Polymorphic variation at the BAT-25 and BAT-26 loci in individuals of African origin: implications for microsatellite instability testing. Am J Pathol 155: 349–353.
Rampino N, Yamamoto H, Ionov Y, Li Y, Sawai H, Reed JC et al. (1997). Somatic frameshift mutations in the BAX gene in colon cancers of the microsatellite mutator phenotype. Science 275: 967–969.
Ribic CM, Sargent DJ, Moore MJ, Thibodeau SN, French AJ, Goldberg RM et al. (2003). Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N Engl J Med 349: 247–257.
Rodriguez-Bigas M, Boland C, Hamilton S, Henson D, Jass J, Khan P et al. (1997). A national cancer institute workshop on hereditary nonpolyposis colorectal cancer syndrome: meeting highlights and Bethesda guidelines. J Natl Cancer Inst 89: 1758–1762.
Roncalli M, Bianchi P, Grimaldi G, Ricci D, Laghi L, Maggioni M et al. (2000). Fractional allelic loss in non-end-stage cirrhosis: correlations with hepatocellular carcinoma development during follow-up. Hepatology 31: 846–850.
Samowitz W, Albertsen H, Herrick J, Levin T, Sweeney C, Murtaugh M et al. (2005). Evaluation of a large, population-based sample supports a CpG island methylator phenotype in colon cancer. Gastroenterology 129: 837–845.
Samowitz WS, Curtin K, Ma K-N, Schaffer D, Coleman LW, Leppert M et al. (2001). Microsatellite instability in sporadic colon cancer is associated with an improved prognosis at the population level. Cancer Epidemiol Biomarkers Prev 10: 917–923.
Strachan TR . (1999). Human Molecular Genetics 2. John Wiley & Sons: New York.
Suraweera N, Duval A, Reperant M, Vaury C, Furlan D, Leroy K et al. (2002). Evaluation of tumor microsatellite instability using five quasimonomorphic mononucleotide repeats and pentaplex PCR. Gastroenterology 123: 1804–1811.
Thibodeau S, Bren G, Schaid D . (1993). Microsatellite instability in cancer of the proximal colon. Science 260: 816–819.
Thibodeau SN, French AJ, Cunningham JM, Tester D, Burgart LJ, Roche PC et al. (1998). Microsatellite instability in colorectal cancer: different mutator phenotypes and the principal involvement of hMLH1. Cancer Res 58: 1713–1718.
Thibodeau SN, French AJ, Roche PC, Cunningham JM, Tester DJ, Lindor NM et al. (1996). Altered expression of hMSH2 and hMLH1 in tumors with microsatellite instability and genetic alterations in mismatch repair genes. Cancer Res 56: 4836–4840.
Tomlinson I, Halford S, Aaltonen L, Hawkins N, Ward R . (2002). Does MSI-low exist? J Pathol 197: 6–13.
Toyota M, Ahuja N, Ohe-Toyota M, Herman JG, Baylin SB, Issa J-PJ . (1999). CpG island methylator phenotype in colorectal cancer. Proc Natl Acad Sci 96: 8681–8686.
Truninger K, Menigatti M, Luz J, Russell A, Haider R, Gebbers J et al. (2005). Immunohistochemical analysis reveals high frequency of PMS2 defects in colorectal cancer. Gastroenterology 128: 1160–1171.
Umar A, Boland CR, Terdiman JP, Syngal S, Chapelle Adl, Ruschoff J et al. (2004). Revised Bethesda guidelines for hereditary nonpolyposis colorectal cancer (Lynch syndrome) and microsatellite instability. J Natl Cancer Inst 96: 261–268.
Vasen H, Mecklin J, Khan P, Lynch H . (1991). The international collaborative group on hereditary non-polyposis colorectal cancer (ICG-HNPCC). Dis Colon Rectum 34: 424–425.
Watanabe T, Wu T-T, Catalano PJ, Ueki T, Satriano R, Haller DG et al. (2001). Molecular predictors of survival after adjuvant chemotherapy for colon cancer. N Engl J Med 344: 1196–1206.
Weinberg R . (2006). The Biology of Cancer. Garland Science: New York.
Wu Y, Nystrom-Lahti M, Osinga J, Looman M, Peltomaki P, Aaltonen L et al. (1997). MSH2 and MLH1 mutations in sporadic replication error-positive colorectal carcinoma as assessed by two-dimensional DNA electrophoresis. Genes Chromosomes Cancer 18: 269–278.
Xicola RM, Llor X, Pons E, Castells A, Alenda C, Pinol V et al. (2007). Performance of different microsatellite marker panels for detection of mismatch repair-deficient colorectal tumors. J Natl Cancer Inst 99: 244–252.
Yamamoto H, Sawai H, Perucho M . (1997). Frameshift somatic mutations in gastrointestinal cancer of the microsatellite mutator phenotype. Cancer Res 57: 4420–4426.
Yamashita K, Dai T, Dai Y, Yamamoto F, Perucho M . (2003). Genetics supersedes epigenetics in colon cancer phenotype. Cancer Cell 4: 121–131.
Zhou X, Hoang J, Cottu P, Thomas G, Hamelin R . (1997). Allelic profiles of mononucleotide repeat microsatellites in control individuals and in colorectal tumors with and without replication errors. Oncogene 15: 1713–1718.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Laghi, L., Bianchi, P. & Malesci, A. Differences and evolution of the methods for the assessment of microsatellite instability. Oncogene 27, 6313–6321 (2008). https://doi.org/10.1038/onc.2008.217
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2008.217
Keywords
This article is cited by
-
The clinicopathological characteristics, prognosis and immune microenvironment mapping in MSI-H/MMR-D endometrial carcinomas
Discover Oncology (2022)
-
Worldwide variation in lynch syndrome screening: case for universal screening in low colorectal cancer prevalence areas
Familial Cancer (2021)
-
Accuracy of four mononucleotide-repeat markers for the identification of DNA mismatch-repair deficiency in solid tumors
Journal of Translational Medicine (2018)
-
Evolving notions on immune response in colorectal cancer and their implications for biomarker development
Inflammation Research (2018)
-
Epstein–Barr virus positivity, not mismatch repair-deficiency, is a favorable risk factor for lymph node metastasis in submucosa-invasive early gastric cancer
Gastric Cancer (2016)