Abstract
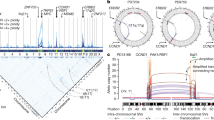
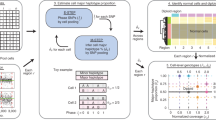
Chromosome translocations in the common epithelial cancers are abundant, yet little is known about them. They have been thought to be almost all unbalanced and therefore dismissed as mostly mediating tumour suppressor loss. We present a comprehensive analysis by array painting of the chromosome translocations of breast cancer cell lines HCC1806, HCC1187 and ZR-75-30. In array painting, chromosomes are isolated by flow cytometry, amplified and hybridized to DNA microarrays. A total of 200 breakpoints were identified and all were mapped to 1 Mb resolution on bacterial artificial chromosome (BAC) arrays, then 40 selected breakpoints, including all balanced breakpoints, were further mapped on tiling-path BAC arrays or to around 2 kb resolution using oligonucleotide arrays. Many more of the translocations were balanced at 1 Mb resolution than expected, either reciprocal (eight in total) or balanced for at least one participating chromosome (19 paired breakpoints). Second, many of the breakpoints were at genes that are plausible targets of oncogenic translocation, including balanced breaks at CTCF, EP300/p300 and FOXP4. Two gene fusions were demonstrated, TAX1BP1–AHCY and RIF1–PKD1L1. Our results support the idea that chromosome rearrangements may play an important role in common epithelial cancers such as breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Abdel-Rahman WM, Katsura K, Rens W, Gorman PA, Sheer D, Bicknell D et al. (2001). Spectral karyotyping suggests additional subsets of colorectal cancers characterized by pattern of chromosome rearrangement. Proc Natl Acad Sci USA 98: 2538–2543.
Adeyinka A, Kytola S, Mertens F, Pandis N, Larsson C . (2000). Spectral karyotyping and chromosome banding studies of primary breast carcinomas and their lymph node metastases. Int J Mol Med 5: 235–240.
Alsop AE, Teschendorff AE, Edwards PA . (2006). Distribution of breakpoints on chromosome 18 in breast, colorectal, and pancreatic carcinoma cell lines. Cancer Genet Cytogenet 164: 97–109.
Barlund M, Monni O, Weaver JD, Kauraniemi P, Sauter G, Heiskanen M et al. (2002). Cloning of BCAS3 (17q23) and BCAS4 (20q13) genes that undergo amplification, overexpression, and fusion in breast cancer. Genes Chromosomes Cancer 35: 311–317.
Briand P, Petersen OW, Van Deurs B . (1987). A new diploid nontumorigenic human breast epithelial cell line isolated and propagated in chemically defined medium. In vitro Cell Dev Biol 23: 181–188.
Carlsson P, Mahlapuu M . (2002). Forkhead transcription factors: key players in development and metabolism. Dev Biol 250: 1–23.
Chaffanet M, Gressin L, Preudhomme C, Soenen-Cornu V, Birnbaum D, Pebusque MJ . (2000). MOZ is fused to p300 in an acute monocytic leukemia with t(8;22). Genes Chromosomes Cancer 28: 138–144.
Chin KT, Chun AC, Ching YP, Jeang KT, Jin DY . (2007). Human T-cell leukemia virus oncoprotein tax represses nuclear receptor-dependent transcription by targeting coactivator TAX1BP1. Cancer Res 67: 1072–1081.
Davidson JM, Gorringe KL, Chin SF, Orsetti B, Besret C, Courtay-Cahen C et al. (2000). Molecular cytogenetic analysis of breast cancer cell lines. Br J Cancer 83: 1309–1317.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S et al. (2002). Mutations of the BRAF gene in human cancer. Nature 417: 949–954.
Delmas P . (2005). Polycystins: polymodal receptor/ion-channel cellular sensors. Pflugers Arch 451: 264–276.
Dorssers LC, Veldscholte J . (1997). Identification of a novel breast-cancer-anti-estrogen-resistance (BCAR2) locus by cell-fusion-mediated gene transfer in human breast-cancer cells. Int J Cancer 72: 700–705.
Dutrillaux B . (1995). Pathways of chromosome alteration in human epithelial cancers. Adv Cancer Res 67: 59–82.
Engel LW, Young NA, Tralka TS, Lippman ME, O’Brien SJ, Joyce MJ . (1978). Establishment and characterization of three new continuous cell lines derived from human breast carcinomas. Cancer Res 38: 3352–3364.
Fiegler H, Carr P, Douglas EJ, Burford DC, Hunt S, Scott CE et al. (2003a). DNA microarrays for comparative genomic hybridization based on DOP-PCR amplification of BAC and PAC clones. Genes Chromosomes Cancer 36: 361–374.
Fiegler H, Gribble SM, Burford DC, Carr P, Prigmore E, Porter KM et al. (2003b). Array painting: a method for the rapid analysis of aberrant chromosomes using DNA microarrays. J Med Genet 40: 664–670.
Gayther SA, Batley SJ, Linger L, Bannister A, Thorpe K, Chin SF et al. (2000). Mutations truncating the EP300 acetylase in human cancers. Nat Genet 24: 300–303.
Gazdar AF, Kurvari V, Virmani A, Gollahon L, Sakaguchi M, Westerfield M et al. (1998). Characterization of paired tumor and non-tumor cell lines established from patients with breast cancer. Int J Cancer 78: 766–774.
Gribble SM, Kalaitzopoulos D, Burford DC, Prigmore E, Selzer RR, Ng BL et al. (2007). Ultra-high resolution array painting facilitates breakpoint sequencing. J Med Genet 44: 51–58.
Gribble SM, Prigmore E, Burford DC, Porter KM, Ng BL, Douglas EJ et al. (2005). The complex nature of constitutional de novo apparently balanced translocations in patients presenting with abnormal phenotypes. J Med Genet 42: 8–16.
Grigorova M, Lyman RC, Caldas C, Edwards PA . (2005). Chromosome abnormalities in 10 lung cancer cell lines of the NCI-H series analyzed with spectral karyotyping. Cancer Genet Cytogenet 162: 1–9.
Hahn Y, Bera TK, Gehlhaus K, Kirsch IR, Pastan IH, Lee B . (2004). Finding fusion genes resulting from chromosome rearrangement by analyzing the expressed sequence databases. Proc Natl Acad Sci USA 101: 13257–13261.
Huang AL, Chen X, Hoon MA, Chandrashekar J, Guo W, Trankner D et al. (2006). The cells and logic for mammalian sour taste detection. Nature 442: 934–938.
Huang HE, Chin SF, Ginestier C, Bardou VJ, Adelaide J, Iyer NG et al. (2004). A recurrent chromosome breakpoint in breast cancer at the NRG1/neuregulin 1/heregulin gene. Cancer Res 64: 6840–6844.
Ichimura K, Mungall AJ, Fiegler H, Pearson DM, Dunham I, Carter NP et al. (2006). Small regions of overlapping deletions on 6q26 in human astrocytic tumours identified using chromosome 6 tile path array-CGH. Oncogene 25: 1261–1271.
Ida K, Kitabayashi I, Taki T, Taniwaki M, Noro K, Yamamoto M et al. (1997). Adenoviral E1A-associated protein p300 is involved in acute myeloid leukemia with t(11;22)(q23;q13). Blood 90: 4699–4704.
Krzywinski M, Bosdet I, Smailus D, Chiu R, Mathewson C, Wye N et al. (2004). A set of BAC clones spanning the human genome. Nucleic Acids Res 32: 3651–3660.
Liu X, Baker E, Eyre HJ, Sutherland GR, Zhou M. . (1999). Gamma-heregulin: a fusion gene of DOC-4 and neuregulin-1 derived from a chromosome translocation. Oncogene 18: 7110–7114.
McMullan TW, Crolla JA, Gregory SG, Carter NP, Cooper RA, Howell GR et al. (2002). A candidate gene for congenital bilateral isolated ptosis identified by molecular analysis of a de novo balanced translocation. Hum Genet 110: 244–250.
Mehra R, Tomlins SA, Shen R, Nadeem O, Wang L, Wei JT et al. (2007). Comprehensive assessment of TMPRSS2 and ETS family gene aberrations in clinically localized prostate cancer. Mod Pathol 20: 538–544.
Mitelman F . (2000). Recurrent chromosome aberrations in cancer. Mutat Res 462: 247–253.
Mitelman F, Johansson B, Mertens F . (2004). Fusion genes and rearranged genes as a linear function of chromosome aberrations in cancer. Nat Genet 36: 331–334.
Mitelman F, Mertens F, Johansson B . (2005). Prevalence estimates of recurrent balanced cytogenetic aberrations and gene fusions in unselected patients with neoplastic disorders. Genes Chromosomes Cancer 43: 350–366.
Morris JS, Carter NP, Ferguson-Smith MA, Edwards PA . (1997). Cytogenetic analysis of three breast carcinoma cell lines using reverse chromosome painting. Genes Chromosomes Cancer 20: 120–139.
Murrell A, Heeson S, Reik W . (2004). Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nat Genet 36: 889–893.
Ng BL, Carter NP . (2006). Factors affecting flow karyotype resolution. Cytometry A 69: 1028–1036.
Ogryzko VV, Schiltz RL, Russanova V, Howard BH, Nakatani Y . (1996). The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell 87: 953–959.
Persson K, Pandis N, Mertens F, Borg A, Baldetorp B, Killander D et al. (1999). Chromosomal aberrations in breast cancer: a comparison between cytogenetics and comparative genomic hybridization. Genes Chromosomes Cancer 25: 115–122.
Pole JC, Courtay-Cahen C, Garcia MJ, Blood KA, Cooke SL, Alsop AE et al. (2006). High-resolution analysis of chromosome rearrangements on 8p in breast, colon and pancreatic cancer reveals a complex pattern of loss, gain and translocation. Oncogene 25: 5693–5706.
Radvanyi L, Singh-Sandhu D, Gallichan S, Lovitt C, Pedyczak A, Mallo G et al. (2005). The gene associated with trichorhinophalangeal syndrome in humans is overexpressed in breast cancer. Proc Natl Acad Sci USA 102: 11005–11010.
Roschke AV, Tonon G, Gehlhaus KS, McTyre N, Bussey KJ, Lababidi S et al. (2003). Karyotypic complexity of the NCI-60 drug-screening panel. Cancer Res 63: 8634–8647.
Rowley JD . (1998). The critical role of chromosome translocations in human leukemias. Annu Rev Genet 32: 495–519.
Seng TJ, Ichimura K, Liu L, Tingby O, Pearson DM, Collins VP . (2005). Complex chromosome 22 rearrangements in astrocytic tumors identified using microsatellite and chromosome 22 tile path array analysis. Genes Chromosomes Cancer 43: 181–193.
Shillingford JM, Murcia NS, Larson CH, Low SH, Hedgepeth R, Brown N et al. (2006). The mTOR pathway is regulated by polycystin-1, and its inhibition reverses renal cystogenesis in polycystic kidney disease. Proc Natl Acad Sci USA 103: 5466–5471.
Silverman J, Takai H, Buonomo SB, Eisenhaber F, de Lange T . (2004). Human Rif1, ortholog of a yeast telomeric protein, is regulated by ATM and 53BP1 and functions in the S-phase checkpoint. Genes Dev 18: 2108–2119.
Sinclair PB, Nacheva EP, Leversha M, Telford N, Chang J, Reid A et al. (2000). Large deletions at the t(9;22) breakpoint are common and may identify a poor-prognosis subgroup of patients with chronic myeloid leukemia. Blood 95: 738–743.
Sirivatanauksorn V, Sirivatanauksorn Y, Gorman PA, Davidson JM, Sheer D, Moore PS et al. (2001). Non-random chromosomal rearrangements in pancreatic cancer cell lines identified by spectral karyotyping. Int J Cancer 91: 350–358.
Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber T et al. (2006). The Consensus Coding Sequences of Human Breast and Colorectal Cancers. Science 314: 268–274.
Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S et al. (2007). Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 448: 561–566.
Theodorou V, Kimm MA, Boer M, Wessels L, Theelen W, Jonkers J et al. (2007). MMTV insertional mutagenesis identifies genes, gene families and pathways involved in mammary cancer. Nat Genet 39: 759–769.
Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW et al. (2005). Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310: 644–648.
Vogelstein B, Kinzler KW . (2004). Cancer genes and the pathways they control. Nat Med 10: 789–799.
Volik S, Zhao S, Chin K, Brebner JH, Herndon DR, Tao Q et al. (2003). End-sequence profiling: sequence-based analysis of aberrant genomes. Proc Natl Acad Sci USA 100: 7696–7701.
Wang XZ, Jolicoeur EM, Conte N, Chaffanet M, Zhang Y, Mozziconacci MJ et al. (1999). gamma-heregulin is the product of a chromosomal translocation fusing the DOC4 and HGL/NRG1 genes in the MDA-MB-175 breast cancer cell line. Oncogene 18: 5718–5721.
Yuasa T, Venugopal B, Weremowicz S, Morton CC, Guo L, Zhou J . (2002). The sequence, expression, and chromosomal localization of a novel polycystic kidney disease 1-like gene, PKD1L1, in human. Genomics 79: 376–386.
Acknowledgements
We thank Mira Grigorova for unpublished SKY data; Elizabeth Batty for help with array analysis; Danita Pearson, Martin McCabe, Wellcome Trust Sanger Institute Microarray Facility, CRUK Microarray Facility, Institute of Cancer Research and University of Cambridge Department of Pathology Microarray Facility for array production. This work was supported by Cancer Research UK; Breast Cancer Campaign, Breast Cancer Research Trust; Samantha Dickson Brain Tumour Trust; Wellcome Trust (NPC, BLN) and studentships from Cambridge Commonwealth Trust and the Sackler Foundation (KAB) and MRC (SLC).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc).
Supplementary information
Rights and permissions
About this article
Cite this article
Howarth, K., Blood, K., Ng, B. et al. Array painting reveals a high frequency of balanced translocations in breast cancer cell lines that break in cancer-relevant genes. Oncogene 27, 3345–3359 (2008). https://doi.org/10.1038/sj.onc.1210993
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210993
Keywords
This article is cited by
-
Ku70 affects the frequency of chromosome translocation in human lymphocytes after radiation and T-cell acute lymphoblastic leukemia
Radiation Oncology (2022)
-
TFEB-VEGFA (6p21.1) co-amplified renal cell carcinoma: a distinct entity with potential implications for clinical management
Modern Pathology (2017)
-
Downregulation of FOXP2 promoter human hepatocellular carcinoma cell invasion
Tumor Biology (2015)
-
Characterization of genetic rearrangements in esophageal squamous carcinoma cell lines by a combination of M-FISH and array-CGH: further confirmation of some split genomic regions in primary tumors
BMC Cancer (2012)
-
Structural analysis of the genome of breast cancer cell line ZR-75-30 identifies twelve expressed fusion genes
BMC Genomics (2012)