Abstract
Background
PVT1 was up-regulated in patients with gastric cancer (GC) and might be as a novel biomarker for predicting GC. However, the exact mechanism of PVT1 exerting functions in GC was still poorly understood. Emerging evidence suggests that long noncoding RNAs may act as endogenous microRNA (miRNA) sponges to bind to miRNAs and regulate their function.
Aim
This study aimed to determine the function of PVT1 on miR-152 expression in GC cells.
Methods
The levels of PVT1 and miR-152 were determined in GC tissues by quantitative real-time PCR. The expression of miR-152 was detected in GC cells transfected with PVT1 plasmid or siPVT1. Luciferase assay was performed to verify the regulation of miR-152 to CD151 or FGF2 expression and PVT1 to miR-152 expression. The effects of PVT1 on the expression of CD151 and FGF2 were evaluated by Western blot.
Results
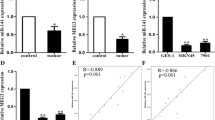
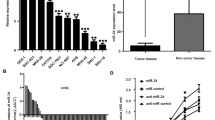
PVT1 was up-regulated in GC tissues than that in the matched normal tissues, and mRNA level of miR-152 was decreased. MiR-152 was negatively associated with PVT1 expression in GC tissues. Based on the in silico analysis, we found that PVT1 have three binding sequences for miR-152. Moreover, PVT1 might inhibit the expression of miR-152 and increased the expression of CD151 and FGF2 through regulating miR-152. PVT1 was positively associated with CD151 and FGF2 expression in GC tissues.
Conclusions
PVT1 might act as a “sponge” to inhibit miR-152 in gastric cancer cells. PVT1 is a promising molecular target to improve the diagnosis and therapy of GC.




Similar content being viewed by others
References
Shin H-R, Carlos MC, Varghese C. Cancer control in the Asia Pacific region: current status and concerns. Jpn J Clin Oncol. 2012;42:867–881.
Crew K-D, Neugut A-I. Epidemiology of gastric cancer. Semin Oncol. 1996;23:281–291.
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA: A Cancer J Clin. 2011;61:69–90.
Wang Y, Feng X, Jia R, et al. Microarray expression profile analysis of long non-coding RNAs of advanced stage human gastric cardia adenocarcinoma. Mol Genet Genomics. 2014;289:291–302.
Xu Z-Y, Yu Q-M, Du Y-A, et al. Knockdown of long non-coding RNA HOTAIR suppresses tumor invasion and reverses epithelial–mesenchymal transition in gastric cancer. Int J Biol Sci. 2013;9:587–597.
Cao W-J, Wu H-L, He B-S, Zhang Y-S, Zhang Z-Y. Analysis of long non-coding RNA expression profiles in gastric cancer. World J Gastroenterol. 2013;19:3658–3664.
Ding J, Li D, Gong M, et al. Expression and clinical significance of the long non-coding RNA PVT1 in human gastric cancer. OncoTargets Therapy. 2014;7:1625–1630.
You L, Chang D, Du H-Z, Zhao Y-P. Genome-wide screen identifies PVT1 as a regulator of Gemcitabine sensitivity in human pancreatic cancer cells. Biochem Biophys Res Commun. 2011;407:1–6.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–233.
Wang X, Yu H, Lu X, Zhang P, Wang M, Hu Y. miR-22 suppresses the proliferation and invasion of gastric cancer cells by inhibiting CD151. Biochem Biophys Res Commun. 2014;445:175–179.
Li C, Li JF, Cai Q, et al. MiRNA-199a-3p in plasma as a potential diagnostic biomarker for gastric cancer. Ann Surg Oncol. 2013;20:397–405.
Xu X, Yang X, Xing C, Zhang S, Cao J. miRNA: The nemesis of gastric cancer. Oncol Lett. 2013;6:631–641.
Shin VY, Chu K-M. MiRNA as potential biomarkers and therapeutic targets for gastric cancer. World J Gastroenterol. 2014;20:10432–10439.
Chen Y, Song Y, Wang Z, et al. Altered expression of MiR-148a and MiR-152 in gastrointestinal cancers and its clinical significance. J Gastrointest Surg. 2010;14:1170–1179.
Zhai R, Kan X, Wang B, et al. miR-152 suppresses gastric cancer cell proliferation and motility by targeting CD151. Tumor Biol. 2014;35:11367–11373.
Cheng Z, Ma R, Tan W, Zhang L. MiR-152 suppresses the proliferation and invasion of NSCLC cells by inhibiting FGF2. Exp Mol Med. 2014;46:e112.
Carramusa L, Contino F, Ferro A, et al. The PVT-1 oncogene is a Myc protein target that is overexpressed in transformed cells. J Cell Physiol. 2007;213:511–518.
Barsotti AM, Beckerman R, Laptenko O, Huppi K, Caplen NJ, Prives C. p53-Dependent induction of PVT1 and miR-1204. J Biol Chem. 2012;287:2509–2519.
Takahashi Y, Sawada G, Kurashige J, et al. Amplification of PVT-1 is involved in poor prognosis via apoptosis inhibition in colorectal cancers. Br J Cancer. 2014;110:164–171.
Yang Y-R, Zang S-Z, Zhong C-L, Li Y-X, Zhao S-S, Feng X-J. Increased expression of the lncRNA PVT1 promotes tumorigenesis in non-small cell lung cancer. Int J Clin Exp Pathol. 2014;7:6929–6935.
Zhou Q, Chen F, Zhao J, et al. Long non-coding RNA PVT1 promotes osteosarcoma development by acting as a molecular sponge to regulate miR-195. Oncotarget. 2016;7:82620–82633.
Yuan CL, Li H, Zhu L, Liu Z, Zhou J, Shu Y. Aberrant expression of long noncoding RNA PVT1 and its diagnostic and prognostic significance in patients with gastric cancer. Neoplasma. 2016;63:442–449.
Wang P, Liu Y-h, Yao Y-l, et al. Long non-coding RNA CASC2 suppresses malignancy in human gliomas by miR-21. Cell Signal. 2015;27:275–282.
Li B, Xie Z, Li B. miR-152 functions as a tumor suppressor in colorectal cancer by targeting PIK3R3. Tumor Biol. 2016;37:10075–10084.
Zhu C, Li J, Ding Q, et al. miR-152 controls migration and invasive potential by targeting TGFα in prostate cancer cell lines. Prostate. 2013;73:1082–1089.
Rubinstein E. The complexity of tetraspanins. Biochem Soc Trans. 2011;39:501–505.
Lau M-T, So W-K, Leung PC. Fibroblast growth factor 2 induces E-cadherin down-regulation via PI3K/Akt/mTOR and MAPK/ERK signaling in ovarian cancer cells. PLoS ONE. 2013;8:e59083.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no actual or potential conflicts of interest.
Additional information
Ting Li and Xiang-ling Meng have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, T., Meng, Xl. & Yang, Wq. Long Noncoding RNA PVT1 Acts as a “Sponge” to Inhibit microRNA-152 in Gastric Cancer Cells. Dig Dis Sci 62, 3021–3028 (2017). https://doi.org/10.1007/s10620-017-4508-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-017-4508-z